Georg Steinert
marine ecology and systematics
Microbial communities in sponges (2011 - now)
The main focus of my PhD thesis is the molecular analysis of microbial communities in sponges.
Many marine sponges contain dense and diverse microbial communities of both ecological and biotechnological importance. These micro-organisms, which can comprise up to 35% of total sponge volume, can be affiliated with Eukarya, Archaea and Bacteria, with at least some of these microbes appearing to be largely specific to sponge hosts. Recent high-throughput sequencing studies on sponge-associated microbial communities from different geographic regions and habitats have expanded our knowledge about the microbial composition of low- and high-microbial-abundance sponges, and divided their communities into: 'core', 'variable', and 'species specific' (Schmitt et al. 2012).
Three different transmission scenarios of sponge-specific microorganisms are under consideration: (1) horizontal transmission, (2) vertical transmission, and (3) a combination of both pathways (Taylor et al. 2007).
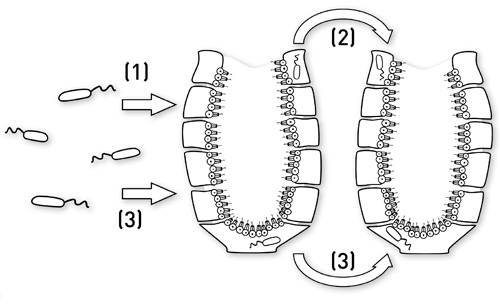
(1) Horizontal transmission
Sponge-specific bacterias are present in low abundance in the surrounding seawater maintained by unspecific or specific enrichment (Taylor et al. 2007):
- specific absorption via recognition (e.g. by the immune system)
- subtractive enrichment - accumulation of phagocystosis resisting bacterias
- unspecific enrichment via filtering of surrounding seawater
(2) Vertical transmission
- sponge-specific phylogenetic cluster from distantly related, geographically separated sponges
- presumably maintained via precambrian acquistion or recent colonization (Taylor et al. 2007)
- sponge-specific bacteria are not present in the surrounding seawater
- results from standard molecular methods (i.e. DGGE, FISH, 16S rRNA sequencing) supported this evolutionary pathway for bacteria associated with sponges
- only one example of symbiont transmission in sponge sperm known - from a cyanobacteria and the Australian sponge Chondrilla australiensis (Webster et al. 2012)
(3) Vertical and environmental transmission
- Vertical transmission is supplemented by environmental aqcuisition of low abundance sponge-specific bacterias from the surrounding seawater
Our ISME14 poster for further informations - click to enlarge:

Diffusion-growth-chambers in sponges
Sponge-microbe associations are renowned as a source of bioactive metabolites, with >290 novel compounds isolated from sponges each year (Taylor et al. 2007). Given that at least some of these metabolites are believed to be made by symbiotic microbes, cultivation of sponge-associated bacteria has been a major focus in the search for novel marine natural products.
However, most bacteria from environmental samples, including sponges, have resisted cultivation on artificial growth media. One promising approach to isolating so-called uncultivable microorganisms is the diffusion chamber method (Bollmann et al. 2007; Kaeberlein et al. 2002).
While there are different variations on this approach, the unifying feature is an attempt to cultivate bacteria in situ by allowing interaction with other microorganisms and chemical components within the ambient environment.
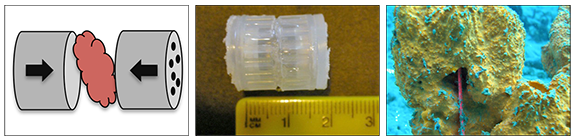
This was the first attempt at using diffusion chambers for the isolation of uncultivable bacteria from sponges. The study was conducted by implanting diffusion growth chambers (DGC) in the tissue of Rhabdastrella globostellata reef sponges.
From left to right: schematic model of a DGC, the final chamber, and within a sponge.

The poster we presented at the YouMares conference 2013 - you can find more information in our Marine Biotechnology publication: 'Application of Diffusion Growth Chambers for the Cultivation of Marine Sponge-Associated Bacteria' - click to enlarge:

Local spatial patterns in sponge-microbe communities
In this study we examined whether abiotic conditions affect the intraspecific microbial communities of sponges. We hypothesized that sponges could act as microbial refugia to sustain disturbances as part of ecosystem resilience. In total, 23 marine sponge specimens (from three species) were collected from two depths (shallow sublittoral, 5-10m and mesophotic zone, 100-120m) around Guam.
Preliminary results suggest that the sponge-associated communities are host-specific (dotted circles). Next step is to analyse the in-host variability (color gradient).
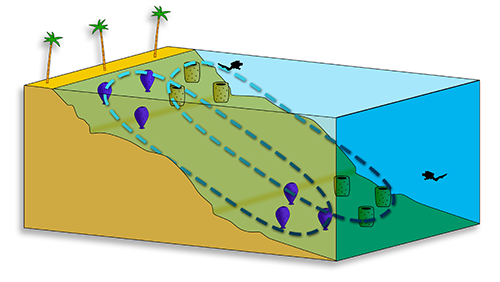
We analyzed the microbial communities within these samples using 16S rRNA gene-targeted amplicon pyrosequencing, from the high-microbial-abundance sponge Rhabdastrella globostellata and two low-microbial-abundance sponges Callyspongia and Acanthella. So far our results suggest that the sponge-associated microbial communities are host specific overall.
First results were presented during the 2nd International Symposium on Sponge Microbiology in Baltimore 2014 - click to enlarge:
References
- Bollmann, A., Lewis, K., & Epstein, S. S. (2007). Incubation of environmental samples in a diffusion chamber increases the diversity of recovered isolates. Applied and environmental microbiology, 73(20), 6386 90.
- Kaeberlein, T., Lewis, K., & Epstein, S. S. (2002). Isolating uncultivable microorganisms in pure culture in a simulated natural environment. Science (New York, N.Y.), 296(5570), 1127 9.
- Schmitt S, Tsai P, Bell J, Fromont J, Ilan M, Lindquist N, et al. (2012). Assessing the complex sponge microbiota: core, variable and species-specific bacterial communities in marine sponges. The ISME journal 6:564 76.
- Taylor MW, Radax R, Steger D, Wagner M. (2007). Sponge-associated microorganisms: evolution, ecology, and biotechnological potential. Microbiology and molecular biology reviews MMBR 71:295 347.
- Webster NS, Taylor MW. (2012). Marine sponges and their microbial symbionts: love and other relationships. Environmental microbiology 14:335 46.